Binding mechanism of triclocarban with human serum albumin: Effect on the conformation and activity of the model transport protein - ScienceDirect
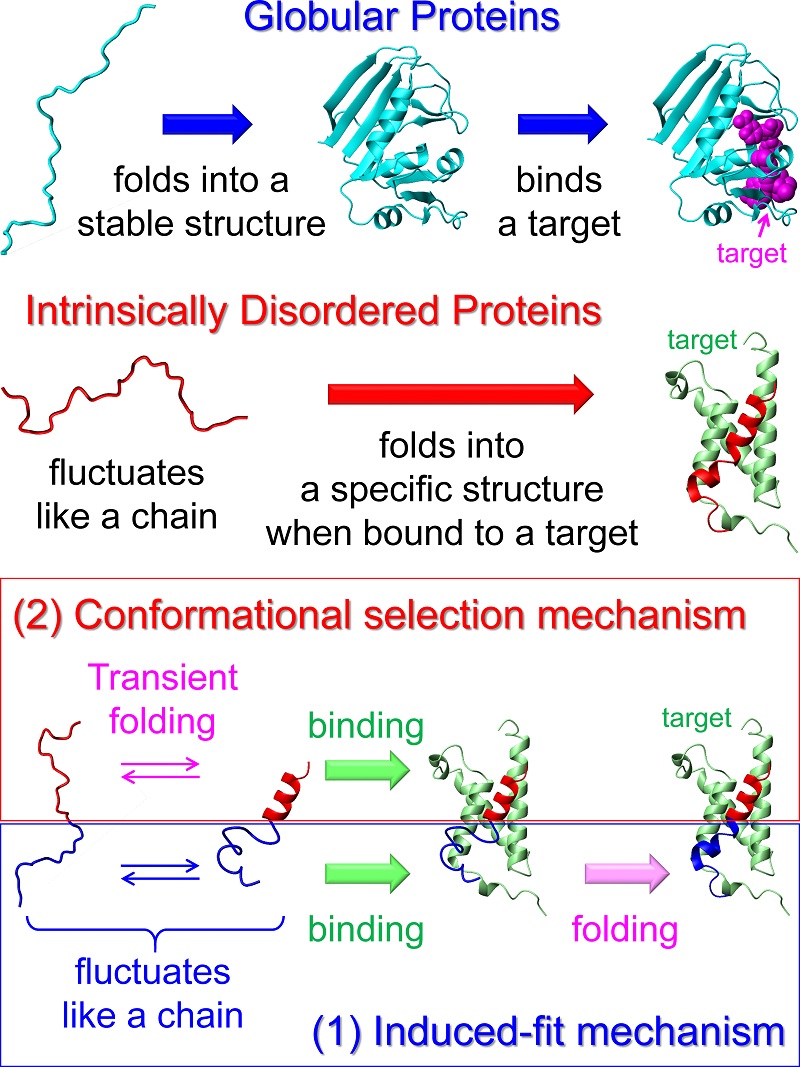
![PDF] Unified understanding of folding and binding mechanisms of globular and intrinsically disordered proteins | Semantic Scholar PDF] Unified understanding of folding and binding mechanisms of globular and intrinsically disordered proteins | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/35a46abf0b8d7690ffa2cd04633796d6b83a0eda/4-Figure2-1.png)
PDF] Unified understanding of folding and binding mechanisms of globular and intrinsically disordered proteins | Semantic Scholar

a Schematic diagram of the MmfR binding mechanism. Binding of MmfR to... | Download Scientific Diagram

Molecular Mechanism of the N501Y Mutation for Enhanced Binding between SARS-CoV-2's Spike Protein and Human ACE2 Receptor | bioRxiv
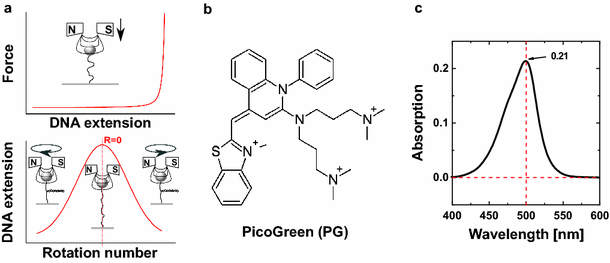
Binding mechanism of PicoGreen to DNA characterized by magnetic tweezers and fluorescence spectroscopy | SpringerLink

The binding mechanism of the virulence factor Streptococcus suis adhesin P subtype to globotetraosylceramide is associated with systemic disease - Journal of Biological Chemistry

Both protein dynamics and ligand concentration can shift the binding mechanism between conformational selection and induced fit | PNAS

Exploring the Binding Mechanism and Accessible Angle of SARS-CoV-2 Spike and ACE2 by Molecular Dynamics Simulation and Free Energy Calculation | Theoretical and Computational Chemistry | ChemRxiv | Cambridge Open Engage
PLOS Computational Biology: Lipidated apolipoprotein E4 structure and its receptor binding mechanism determined by a combined cross-linking coupled to mass spectrometry and molecular dynamics approach
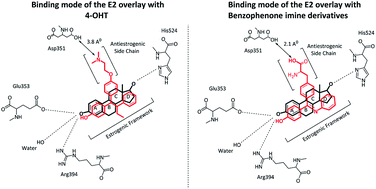
Computational investigations of the binding mechanism of novel benzophenone imine inhibitors for the treatment of breast cancer - RSC Advances (RSC Publishing)

A schematic diagram describing the proposed two-step binding mechanism for proteins in steric occlusion of the direct binding of the ligands.

Insights into the substrate binding mechanism of SULT1A1 through molecular dynamics with excited normal modes simulations | Scientific Reports
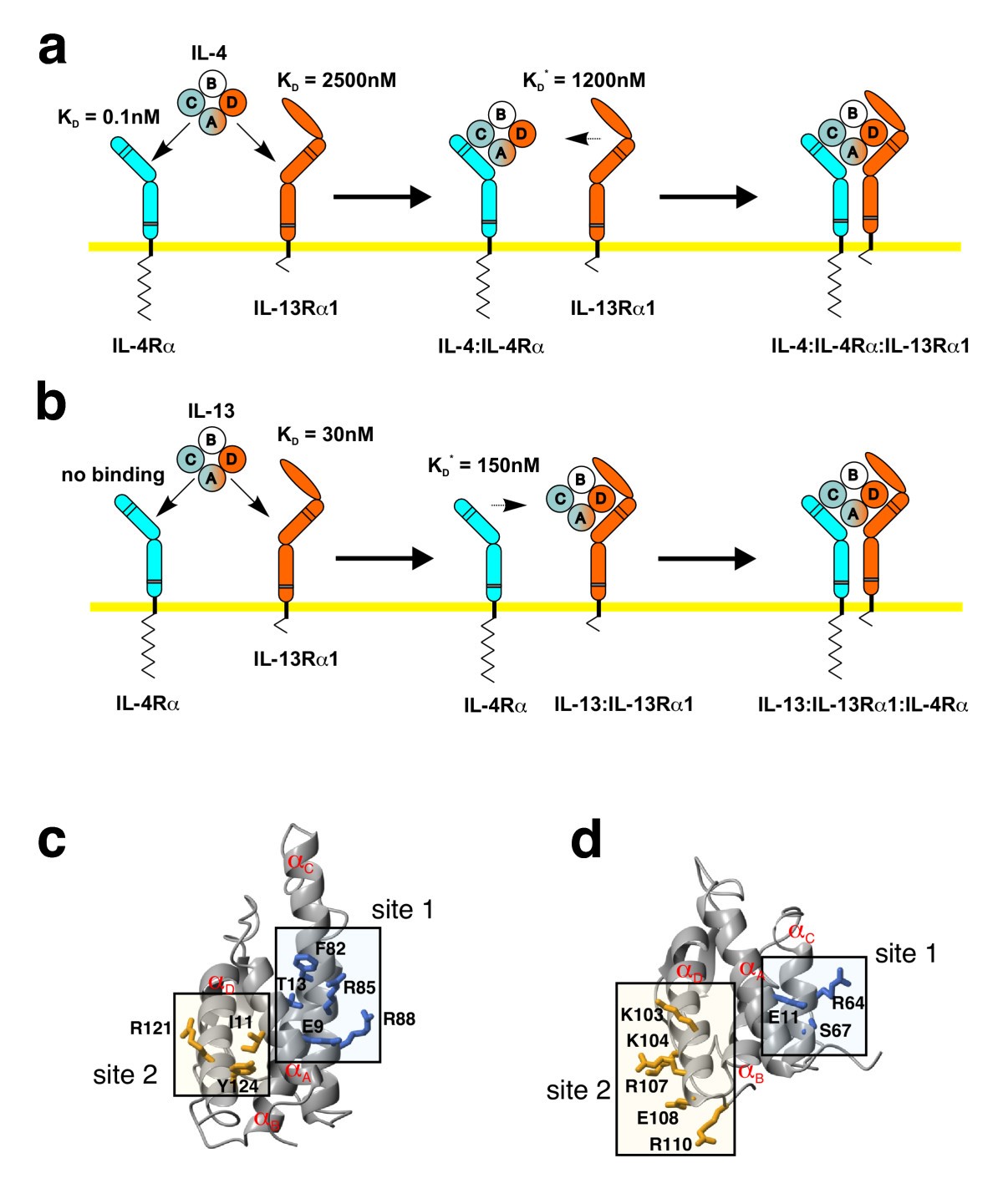
A modular interface of IL-4 allows for scalable affinity without affecting specificity for the IL-4 receptor | BMC Biology | Full Text
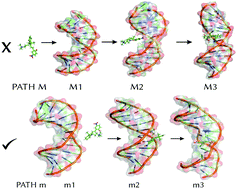
DNA-binding mechanism of spiropyran photoswitches: the role of electrostatics - Physical Chemistry Chemical Physics (RSC Publishing)

Binding mechanism and binding free energy of amino acids and citrate to hydroxyapatite surfaces as a function of crystallographic facet, pH, and electrolytes - ScienceDirect
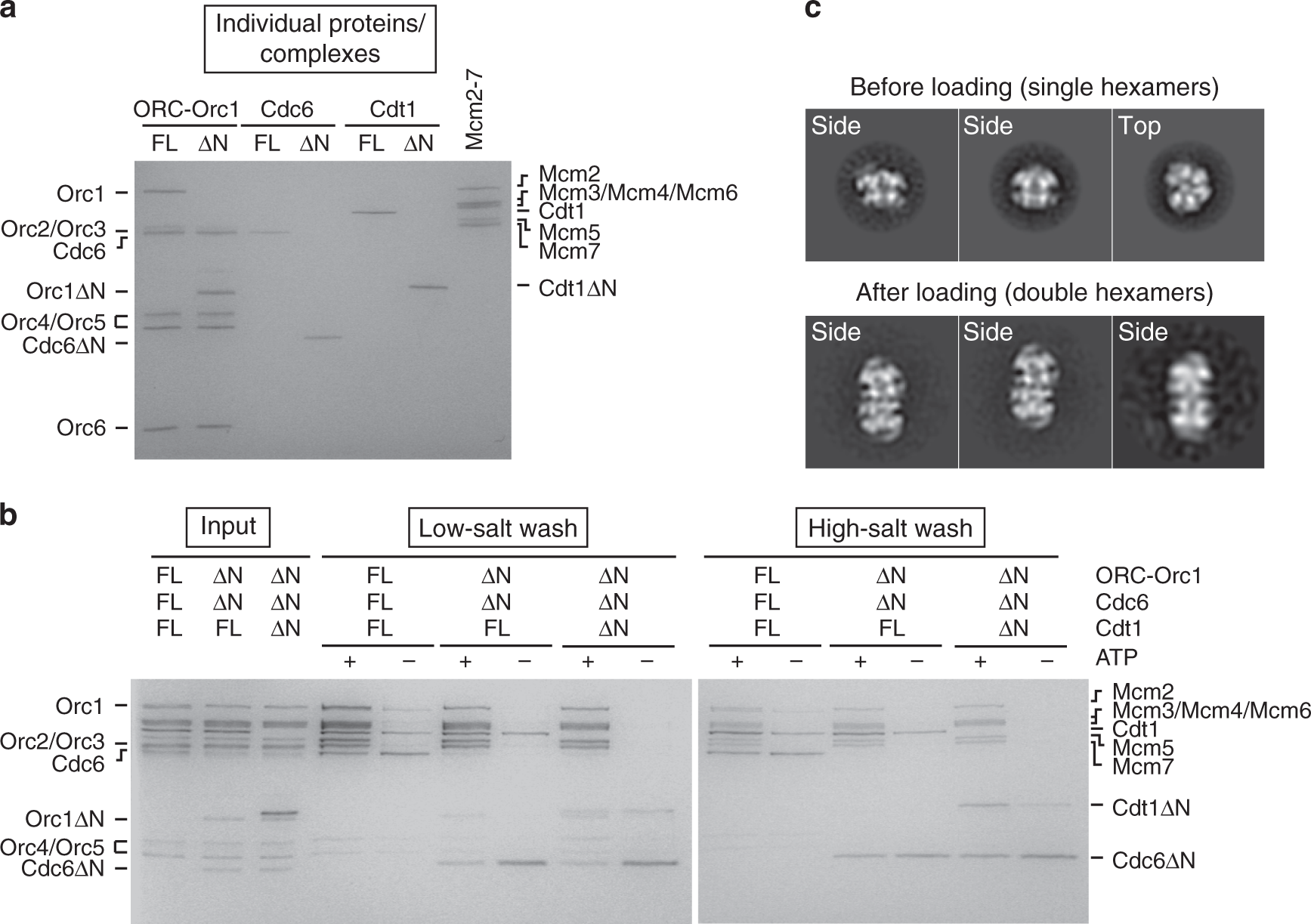
Structural mechanism for replication origin binding and remodeling by a metazoan origin recognition complex and its co-loader Cdc6 | Nature Communications

Identification of the Type I Collagen-binding Domain of Bone Sialoprotein and Characterization of the Mechanism of Interaction* - Journal of Biological Chemistry